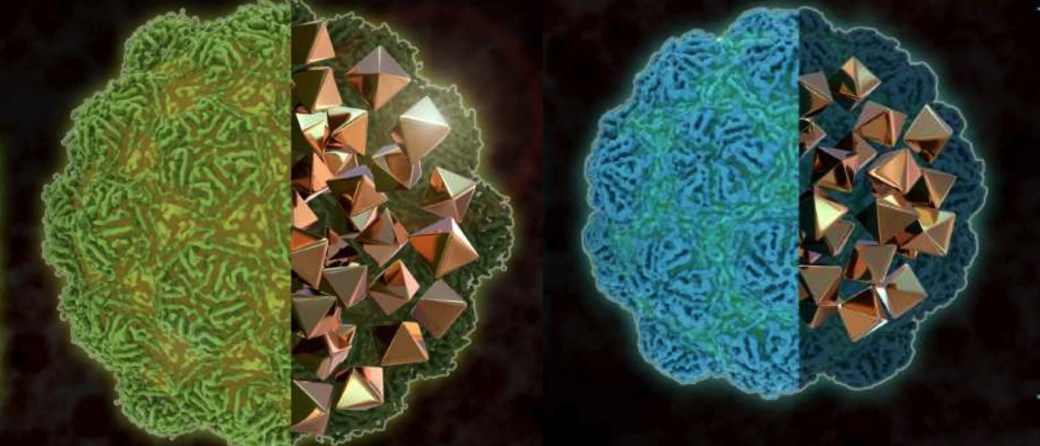
Scientists at the Technical University of Munich (TUM) and at Helmholtz Zentrum München have developed a method to visualize gene expression in cells using electron microscopy. Although electron microscopy provides the most detailed images of cells available today, it usually cannot identify different genetic programs running inside them. The new method presented in the scientific journal ‘ACS Nano’ is based on genetically modified cells that produce nanometer sized spheres with their exact sizes depending on the genetic processes going on in the cell. Thus, by determining the sizes of the spheres, researchers will be able to distinguish between different processes. Furthermore, this method might even help to investigate how memories are stored in neural networks.
What exactly is going on in living cells? This question has kept scientists busy for decades. To label small structures, scientists have been using fluorescent proteins, but these are only visible in light microscopes, which show a relatively poor resolution. In Contrast, electron microscopes provide a more detailed picture, but "so far there are hardly any solutions for multi-color genetic labeling of cells for this technology that would allow to directly tell different cells apart", says Prof. Dr. Gil Gregor Westmeyer. He is Professor of Molecular Imaging at TUM School of Medicine, Principal Investigator at TUM’s Munich School of BioEngineering and head of a research group at the Institute for Biological and Medical Imaging (IBMI) of Helmholtz Zentrum München.
Nanocompartments as multi-Color labels for electron microscopy
Westmeyer and his colleagues have been working with so-called encapsulins for some time. These are small, non-toxic proteins that can usually be found in bacteria. Encapsulins automatically assemble to form nanocompartments, in which selected chemical reactions can take place without disturbing the metabolism of the cell. The scientists have produced genetically modified cells that form nanocompartments with diameters depending on the experimental conditions. "Like the palette of colors in fluorescence microscopy, our method turns geometry into a label for electron microscopy," adds Felix Sigmund from Westmeyer's research group.
To achieve strong contrast in the images produced by the electron microscope, the researchers use the enzyme Ferroxidase, which can be placed inside the encapsulin compartments. When iron ions enter the compartment through its pores, the enzyme oxidizes divalent iron ions into their trivalent form. This creates insoluble iron oxides that remain inside the nanocompartment. Metals provide good contrasts in electron microscopy because they absorb electrons – this can be compared with bones in the human body, which strongly absorb X-rays and thus can be easily distinguished in an X-ray image.
Following neural tracts in the X-ray Image
In a next step, the researchers want to use their new method to investigate neural circuits. Despite the impressive resolution of electron microscopy, the method still cannot reliably distinguish certain types of neurons within the brain. "With our new reporter genes, we might be able to label specific cells and then observe the types of connections made by different nerve cell and determine the cells’ states," adds Westmeyer. This new technology could thus help to find the exact circuit layouts of the brains in different species and to study how memories are stored in neural Networks.
This article is based on a release published by the Helmholtz Zentrum München.
Publication:
Sigmund, Pettinger, Jube et al., “Iron-sequestering nanocompartments as multiplexed Electron Microscopy gene reporters”, ACS Nano, 2019 13 (7), 8114-8123 DOI: 10.1021/acsnano.9b03140
More information:
Short Profile of Prof. Gil Westmeyer: http://www.professoren.tum.de/en/westmeyer-gil/
Website of Gil Westmeyer’s research Group: https://www.westmeyerlab.org
An earlier paper from Gil Westmeyer’s research group reporting on the creation of encapsulin nanocompartments in mammalian cells:
Felix Sigmund et al., “Bacterial encapsulins as orthogonal compartments for mammalian cell engineering”, Nature Communications, Volume 9, Article number: 1990 (2018), 18 May 2018; DOI: 10.1038/s41467-018-04227
Scientific Contact:
Prof. Gil Gregor Westmeyer
Technical University of Munich
Professor of Molecular Imaging
Phone: +49 89 3187 2123
Email: gil.westmeyer@tum.de
Media Relations MSB:
Dr. Paul Piwnicki
Phone: +49 89 289 10808
Email: paul.piwnicki@tum.de